“We acknowledge that you discuss control strategy for the impurities which could be present in the drug substance in sections 3.2.S.2.3, 2.4, and 3.2. We also acknowledged that you have provided risk assessment of potential genotoxicity for the PMIs listed in section 3.2.S.3.2. However, we were unable to locate a hazard assessment for most of the actual as well as potential impurities in your submission. A complete hazard assessment for actual and potential impurities is recommended in Section 6 of the ICH M7 Guidance: Assessment and Control of DNA Reactive (Mutagenic) Impurities in Pharmaceuticals to Limit Potential Carcinogenic Risk. Section 6 of the guidance describes the elements required for impurity hazard assessment and requires classification of each impurity according to Table 1. Classification of all impurities (Classes 1-5) should be justified by either predictive or empirical data and not by visual inspection or statements alone. In addition, please provide (Q)SAR data for the impurities such as Impurity SM2-1, Impurity SM2-2, Impurity SM2-5, Impurity ZJT1-1, Impurity ZJT1-3, Impurity ZJT1-5, Impurity ZJT2-B, and Impurity C to support your proposed classification/justification. Supporting data, as applicable, should accompany the control strategy based on the outcome of the (Q)SAR prediction.”
Based on prior experience in researching mutagenic impurities and the current application, we will discuss the identification and classification of mutagenic impurities.
Previously, in the article "Classification of Carcinogens According to ICH M7", we briefly introduced the classification of carcinogens, which included the identification of mutagenic impurities. The early version of ICH M7 used the term "Genotoxic impurities" for description, but it was later revised to "Mutagenic impurities." This revision indicates that mutagenic substances and genotoxic substances are distinct. Their differences and connections with carcinogens can be illustrated in Figure 1.
A carcinogen is a substance that causes uncontrolled cell proliferation and leads to cancer. Genotoxicity refers to substances that damage the genetic material of cells directly or indirectly. Mutagenic substances are DNA-reactive substances that can cause DNA damage and mutations even at low levels, potentially leading to cancer. These can be identified using standard bacterial reverse mutation tests (Ames test).
All substances can be categorized as either carcinogenic or non-carcinogenic. Mutagenic substances are always genotoxic, but if a mutagenic substance shows no carcinogenicity in a two-year rodent bioassay, it is not classified as a carcinogen. Similarly, some genotoxic substances that are non-mutagenic may also not be carcinogens based on such bioassays.
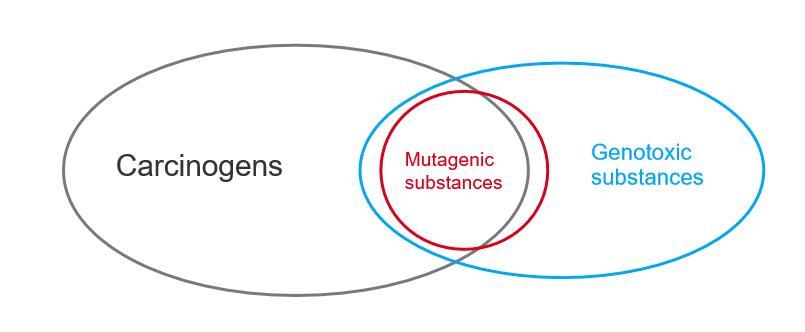
Figure 1. The relationship between carcinogens, genotoxic substances, and mutagenic substances
Take formaldehyde (CAS: 50-00-0) as an example. Formaldehyde is found in textiles, cosmetics, water-treated foods, soy products, and indoor materials like plywood and fiberboards. The Ames test for formaldehyde is positive, indicating DNA breakage or cross-linking in bone marrow cells, making it a mutagenic impurity. However, as an endogenous metabolite and a ubiquitous environmental substance, it may have a reasonably high acceptable intake level of 8000 μg/day or 215 ppb (lower value adopted).
An example is methyl bromide (CAS: 74-83-9). The Ames test for methyl bromide is positive, classifying it as a mutagenic substance. However, its carcinogenicity has not been determined, and the WHO classifies it as a Group 3 carcinogen (not classifiable regarding carcinogenicity in humans). General impurity control suffices for this compound. Toxicological data, such as an oral LD50 in rats of 104 mg/kg and a NOEL of 0.27 mg/kg bw in dogs, can be used to set limits.
Genotoxic substances are categorized as mutagenic or causing chromosomal aberrations. Benzene (CAS: 71-43-2) is an example of a non-mutagenic genotoxic carcinogen. Its Ames test is negative, but it causes chromosomal aberrations or embryonic deformities, making it a chromosomal aberration carcinogen. Such substances are not controlled as mutagenic impurities but require carcinogen-focused research and thresholds (e.g., a limit of 2 ppm for benzene).
An example is carvone (CAS: 2244-16-8). This compound contains structural alerts. The Ames test is negative, while the chromosomal aberration test is positive. It has no carcinogenic activity and is classified as Class 5 under ICH M7, controlled as a general impurity.
Hydroxylamine (CAS: 7803-49-8) is an example. Its Ames test is negative, and it causes cancer through non-DNA-reactive mechanisms. Toxicological data, such as a 24-month study in females showing no effects at 0.4 mg/kg bw/day, support its classification.
ICH M7 focuses on mutagenic impurities. First, relevant carcinogenicity and Ames data of impurities are retrieved through literature and database searches. Based on the results, impurities are classified into Classes 1, 2, and 5. If relevant data cannot be found, SAR (Structure-Activity Relationship) evaluation should be performed, and based on the evaluation results, the impurity should be classified into Classes 3, 4, or 5. However, as described in the defect letter above, "Classification of all impurities (Classes 1-5) should be justified by either predictive or empirical data and not by visual inspection or statements alone." The classification of impurities into Classes 1-5 must be substantiated by predictive or empirical data, rather than relying solely on visual inspection or statements.
In the FDA's March 2021 DMF (Drug Master File) training, the Drug Master File and Drug Substance Workshop – FDA CDER SBIA Events, Day 2: ICH M7(R1) – Chemistry and Manufacturing Control (CMC) Perspective on Hazard Assessment – Barbara O. Scott clearly stated the same point: "NOTE: Visual inspection of a compound for structural alerts for classification purposes is not M7 compliance."
There are now many SAR evaluation software tools available. These tools or methods are based on expert rules or statistical rules. The software I use most often is the expert rule-based Toxtree and the statistical rule-based VEGA. If both software tools produce consistent results, no further discussion is necessary. If the results differ, or if one is positive, commercial evaluation software like Derek/Sarah Nexus is used for further assessment, with the commercial software's result being taken as the final conclusion.
For example, in the supplementary study, we evaluated the impurity SM2-1 using various software tools. First, we used Toxtree software, which gave a high-risk result (Figure 2). We then used VEGA software, which predicted a negative result (Figure 3). Since the two software tools gave opposing conclusions, we used the commercial software Derek/Sarah Nexus to predict the result, which was also negative (Figure 4). Based on these predictions, we concluded that the impurity is mutagenic-negative, classified as a Class 5 impurity, and should be controlled as a general impurity.

Figure 2. SM2-1 mutagenicity prediction results (Toxtree software)

Figure 3. SM2-1 mutagenicity prediction results (VEGA software)

Figure 4. SM2-1 mutagenicity prediction results (Derek Sarah Nexus software)
Therefore, we must first clearly define and classify various impurities, then assess structural alerts, consult literature, and conduct QSAR evaluations. If necessary, Ames tests should be performed. This approach helps prevent delays in the review process caused by supplementary studies, and prevents the failure of product quality due to mutagenic impurities being misclassified as general impurities, which could affect the project evaluation.
[1] Assessment and control of DNA reactive(mutagenic) impurities in pharmaceuticals to limit potential carcinogenic risk. ICH
[2] https://cebs.niehs.nih.gov/cebs/study/002-02185-0005-0000-3. NTP
[3] https://cebs.niehs.nih.gov/datasets/search/ames?casrn=74-83-9. NTP
[4] https://echa.europa.eu/registration-dossier/-/registered-dossier/5298/7/6/1. ECHA
[5] https://echa.europa.eu/registration-dossier/-/registered-dossier/16102/7/7/1. ECHA
[6] Q3C(R9):Impurities: Guideline for Residual Solvents. ICH
[7] https://cebs.niehs.nih.gov/datasets/search/ames?casrn=2244- 16-8. NTP
[8] https://echa.europa.eu/registration-dossier/-/registered-dossier/17464/7/7/1.ECHA
[9] https://echa.europa.eu/registration-dossier/-/registered-dossier/16246/7/6/1.ECHA
 |
 |
 |